Deriving Allergy Substance Category from Recursive SNOMED Calls
📅 16 February, 2018
•⏱️5 min read
As part of some work on recording Adverse Reaction Risks, we took the opportunity to look at binding SNOMED CT into the service. We approached the problem is a very linear way: we need a list of "stuff" that could cause a patient to have an allergic or intolerant reaction. There are a myriad of different term sets available, and SNOMED CT itself has its own.
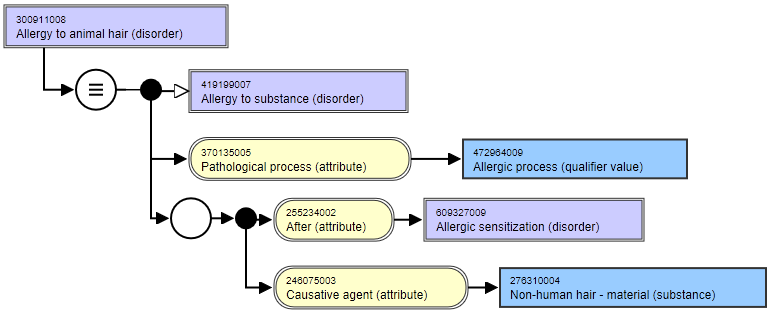
However, by combining a substance with the allergic response you are potentially overly constraining the number of options available to an application. My theory was to record the allergic process separately from the causative agent, and this was backed up by some sterling work by colleagues in the office on patient discharge data. The standard list that SNOMED has of "Allergy to .... " was not big enough.
The other reason for splitting out the agent from the reaction is because it is not always evident that the reaction is indeed allergic; it may be an intolerance or other sensitivity. The difference lies in whether there has been an immunological response to the substance. This will not make a great deal of difference to anyone who is not an immunologist as operational constraints come into play; a ward nurse just needs to know that there could be a significant risk of anaphylaxis if they administer penicillin, and therefore they will avoid that substance and what caused the reaction is moot.
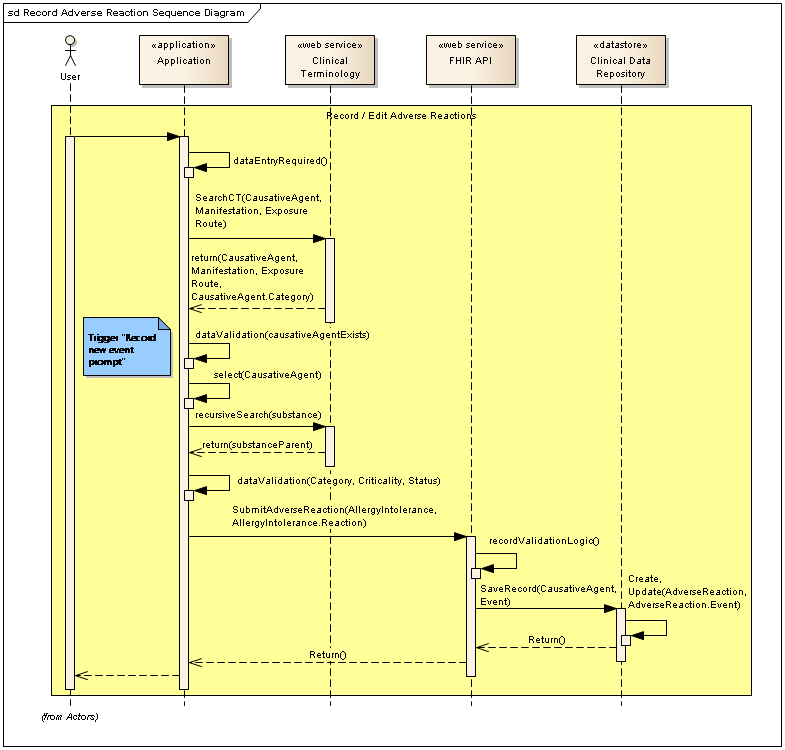
The application will be responsible for providing the user with a search function to interact with the adverse reaction records stored in a repository. We will need to make use of a terminology service, and a FHIR façade to permit search and retrieval functions, something like this...
In the middle of that you can see one of the use cases is to populate the category for the substance as per the HL7 FHIR resource and openEHR "Adverse reaction risk" archetype. In order to populate the FHIR resource AllergyIntolerance.category, a recursive search can be performed once the causative agent is selected by the end user. In this example, the request uses the concept-ID of 10226100 for Strawberry:
GET http://someserver.org.uk:8080/snomedct/concepts/102261002/recursiveParents
(Obviously this is predicated on your Snomed CT service having this capability!) An example edited response:
{
"data": [
{
"id": 72511004,
"conceptId": 72511004,
"fullySpecifiedName": "Fruit (substance)",
"isPrimitive": 1,
"snomedId": "C-F0200"
},
{
"id": 116273005,
"conceptId": 116273005,
"fullySpecifiedName": "Dietary substance (substance)",
"isPrimitive": 1,
"snomedId": "C-F0002"
},
{
"id": 227415002,
"conceptId": 227415002,
"fullySpecifiedName": "Fruit nuts and seeds (substance)",
"isPrimitive": 1,
"snomedId": "C-F068A"
},
**<RESPONSE TRUNCATED>**
{
"id": 227416001,
"conceptId": 227416001,
"fullySpecifiedName": "Fresh fruit (substance)",
"isPrimitive": 1,
"snomedId": "C-F068B"
},
{
"id": 413477004,
"conceptId": 413477004,
"fullySpecifiedName": "Allergen or pseudoallergen (substance)",
"isPrimitive": 1,
"snomedId": "F-61F65"
},
{
"id": 255620007,
"conceptId": 255620007,
"fullySpecifiedName": "Foods (substance)",
"isPrimitive": 1,
"snomedId": "C-F0125"
},
{
"id": 406465008,
"conceptId": 406465008,
"fullySpecifiedName": "Food allergen (substance)",
"isPrimitive": 1,
"snomedId": "F-61DD5"
},
],
"total": n
}With this response and by using the ever-useful Postman, we can then parse the response with some rules to indicate what codes were are looking for. The FHIR specification indicates values of either food, medication, environment or biologic and these can mapped to the SNOMED CT codes indicated in the script. For the causative agent of STRAWBERRY, we want to return the value of Dietary substance. I have used the following SNOMED CT codes to map to FHIR;
115668003 | Biological substance (substance)
116273005 | Dietary substance (substance)
410942007 | Drug or medicament (substance)I am then using environment as a catch all for agents that do not map to the above (which may or may not be the best way to tackle this!).
javascript
pm.test("Concept is medication", function () {
pm.expect(pm.response.text()).to.include("410942007");
});
pm.test("Concept is biological", function() {
pm.expect(pm.response.text()).to.include("115668003");
});
pm.test("Concept is food", function() {
pm.expect(pm.response.text()).to.include("116273005");
});
pm.test("Concept is environmental", function () {
pm.expect(pm.response.text()).to.not.include("115668003");
pm.expect(pm.response.text()).to.not.include("116273005");
pm.expect(pm.response.text()).to.not.include("410942007");
});The order of this logic is important as some causative agents may be listed in more than one category. Owing to the way Snomed CT is structured, you may end up with a food when it is more important to derive medications if you want to reuse the this category for decision support.
Great! And here's the BUT.
Why?!
Why wouldn't we want to do this? In the first instance we are populating the information model and reducing the number of controls presented to the user. That's a good thing.
But this process can be handled later on by real decision support tooling. There is obviously a danger whenever machines use rules to force meaning in the application and this may tread closely to it. Additionally, recent conversations with colleagues seem to indicate that this could be described as meaningless content when you are recording the causative agent as a Snomed term in the first instance. There is an inherent power in capturing good quality, structured data in the first instance that then permits more intelligent use after the event.
While I would generally agree with this, I also think that it does not hurt to include in this case. It's also a good example of how you could use SNOMED-CT to its fullest potential - there's a lot of power under the hood if you unlock it.