Interoperability Badly Needs Defining
📅 13 March, 2018
•⏱️12 min read
It’s not everyday I leaf through my wife’s post to find something to incite the informatics soapbox. But in the most recent issue of the British Journal of General Practice I found just that.
When I usually find myself perusing the BJGP (http://bjgp.org), rarely do I find anything that crosses over into informatics territory. The new issue has the theme “Information for Health” and several interesting articles, with Meaker et al. catching my eye due to the “integration or interoperability” comparison below the title.
The Issue
On first reading, I would wonder how you could conflate the two concepts as if they were an either/or choice. To my mind they are not, but for the authors to reflect on a real world examples work carried out in North East London, there seemed to be a pretty binary theme. They reflected on development of the Vision GP system in piloting models of integrated care. The first was development being commissioned for the currently used Vision system called Health 1000. This used the current application as a foundation and built additional functionality over it. The second was to take a range of operational software from across Primary Care to provide “real-time read/write interoperability”. This involved several systems and revealed three key themes at play; user experience, information governance and the commercial interests of software companies.
I won’t argue with their findings as they make perfect sense. Firstly, users would naturally want to maintain a consistent look and feel to their software and deviations would impact the usability. Secondly, information governance (which is generally the fly in the ointment of electronic communication) provided difficulties for the pilot to negotiate permissions to share data. This resulted in all data being stored on the host practice systems. There was also no master patient index and demographics had to be combined and matched across 137 practices. Not easy work...
But on the third point, the author makes a comment about interoperability relying on the “willingness of software suppliers to work each other”. Playing devils advocate, why would vendors want to work to share data? They are there to make money, not be altruistic. What the author needed to be aware of was that the concept of an open platform needs to be employed here. I can see the difficulty in a pilot to try and get the jigsaw pieces to fit, but that is the purpose of the pilot: to learn. The next step is to take this learning and devise a way of making sure that vendors conform to common standards of data modelling and messaging.
Going back to the beginning of the article, I think I can see where some confusion may stem from. Interoperability is defined by the authors as;
where one software application makes use of data that have been stored in a separate software application and the data transferred have a common meaning, for example, transactions between a bank and an online vendor.
This is one real world example, in very general terms, of an interoperability challenge but it doesn’t clearly delineate between technical, semantic and process. In this scenario, the definition merely reflects the status quo of primary care systems.
I highlighted 'stored' for a reason and I believe that it is wrong to include it here. Assuming the data is stored in the host system is one thing, but that is not to say it cannot be held in some other central clinical repository if the requirements dictate.
I appreciate that this challenge is of particular significance in NHS England, and I do not mean to overly simplify the difficulties that the pilot team had on their hands. However, interoperability is being misused as a term without the basic understanding of the breadth that it encompasses. For anyone to get a full definition, I’d recommend Benson & Grieve (2016) Chapter 2 on “Why Interoperability Is Hard”.
The second definition concerns “integration” where;
a single software system has been developed to cover all activities occurring in an organisation or healthcare system, such as that used by Kaiser Permanente.
Again a quite valid observation and I understand that Kaiser Permanente is not the typical American healthcare provider, being non-profit organisation. However, the usual business model of the large US-based providers is such that controlling each aspect of healthcare is within their gift, and they have the investment to develop solutions meeting these requirements. Therefore it is relatively easy to get a monolithic system to serve your core business needs - they are to all intents within their own walled garden. But the NHS is markedly different in terms of provision across primary, secondary and tertiary care settings, and the author's own assertions of the difficulty in linking the demographics of 137 proves the point. Add in that the 137 practices are a myriad of separate organisations and there is an additional difficulty to overcome.
A Better Definition
The well conceived definition of interoperability derives from IEEE:
Interoperability is ability of two or more systems or components to exchange information and to use the information that has been exchanged (IEEE 1990)
HIMSS define it in a similar way to IEEE but also describe
- foundational; basic data exchange between systems,
- structural; data exchanged has a defined standardised format, and
- semantic; where two "two or more systems or elements to exchange information and to use the information that has been exchanged" (which is also derived from IEEE).
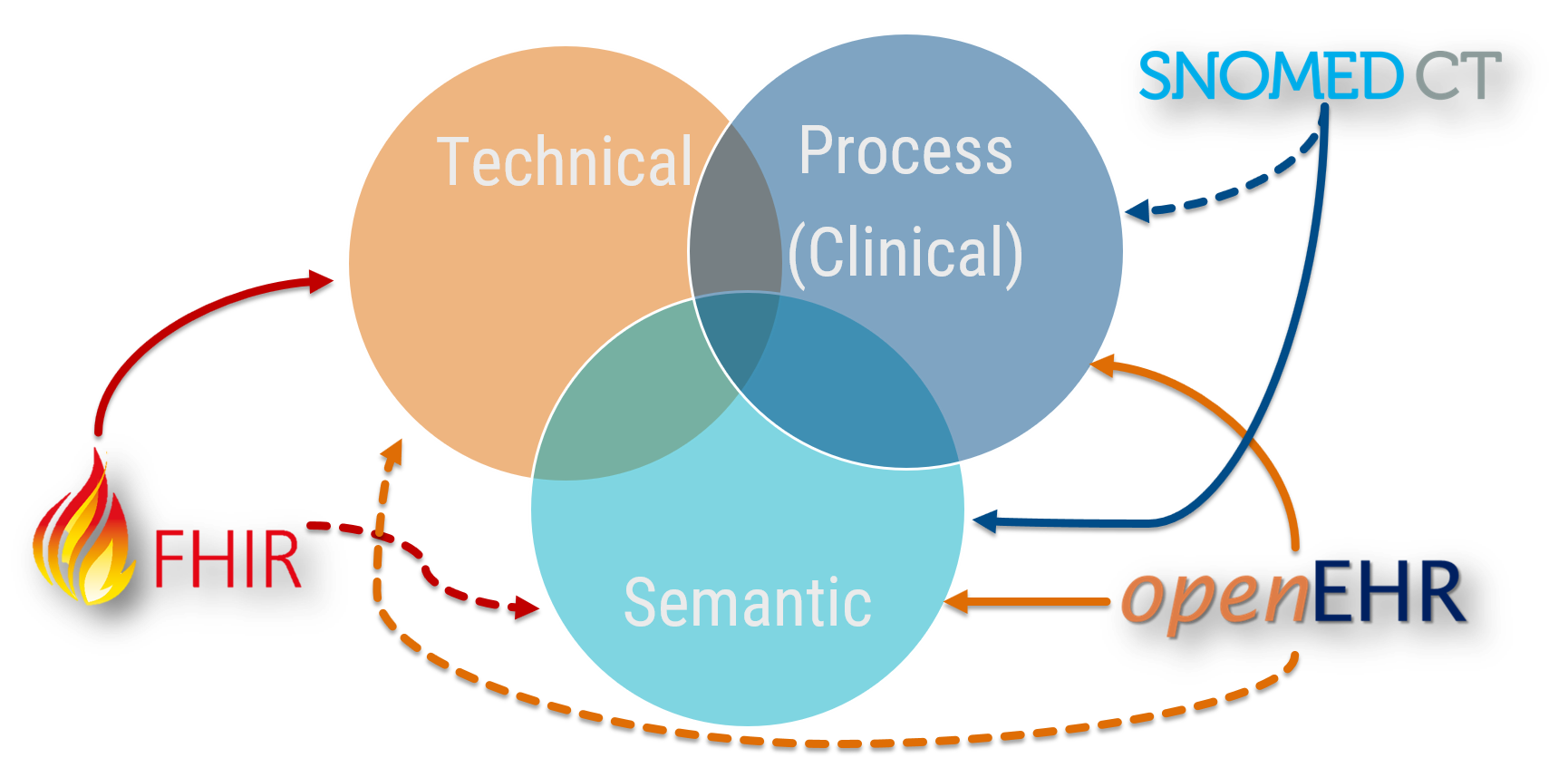
Benson and Grieve also expand upon the basic definition and describe a similar three further facets of granularity (what I have come to know as The Holy Triumvirate of Clinical Data Interoperability):
At any point, clinical data can be described wholly or in part in terms of technical, semantic or process based interoperability. That is not to say that interoperability needs each of these facets at all times, but depending on the use case there will be an inevitable a mixture going on where they intersect in the Reuleaux Triangle1 of interoperable joy. Also, some technologies support these facets better than others, and are indicated by dashed lines.
So to break this down;
- Technical interoperability moves data from one system to another system. Technical interoperability is only concerned about message itself, and not the contents. It is the most solid aspect of interoperability, proven for decades with technologies such as HL7 v2 and more recently with HL7 FHIR.
- Semantic interoperability is achieved when the meaning of data is exchanged and is unambiguously defined. It supports the concept of clinical provenance as it is specific to individual clinical domains and the context in which it has been recorded. This is additionally supported through clinical terminologies, codes and identifiers (such as Snomed CT). This is the principle use case of openEHR.
- Process (i.e. clinical) interoperability is achieved when clinical users are able to share understanding across disparate workflows and business processes. The care context is thus understood as well as the data.
System Engineering Definitions Differ
There is quite some difference between the pure academia descriptions of interoperability and those more commonly used at the coal face, and described above. Grieve's definition of clinical interoperability is;
the ability to transfer patients between care teams and provide seamless provision of clinical care. That is the interoperability that matters and will make a difference to people's lives.
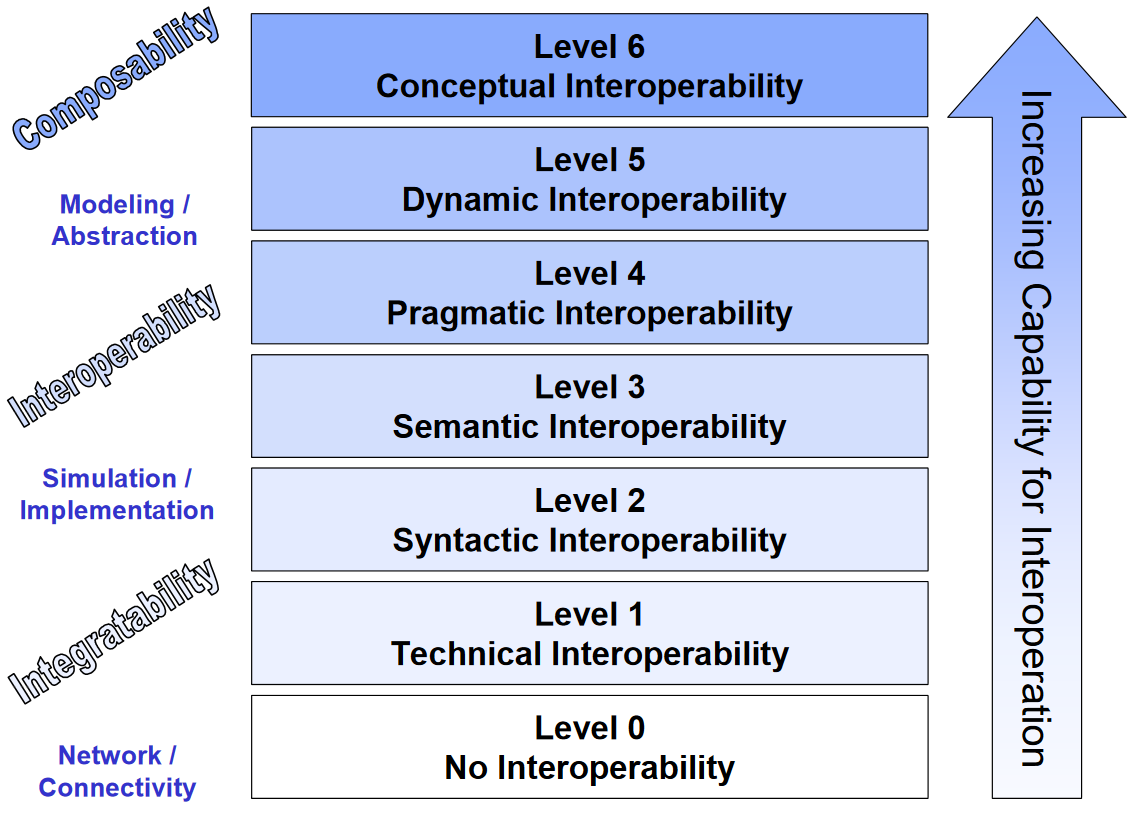
Tolk et al. describe finer granularity in their Levels of Conceptual Interoperability Model (LCIM). The highest levels describes interoperability that becomes situationally aware, and can react accordingly. But essentially, 'process interoperability' as described by Benson and Grieve is an amalgamation of a slightly different facet of semantic interoperability (semantic+?), and Level 4 'pragmatic interoperability' (shown below).
(I'll be looking at this, and specifically syntactic interoperability in the future and no doubt will have something to say on the subject...)
Putting This Into Real Language
Use Case 1: Labs Message
A simple electronic message giving laboratory results for creatinine or HBa1C. When system A sends it, and system B knows what it contains, the message can be received and stored. The structure of the message may contain terminological data, be proprietary or contain non-standard data. The message may contain certain elements to describe its contents and use common language such as XML.
This example describes technical interoperability.
Use Case 2: Labs Message Embedded in a Document
This takes the content of use case 1, and adds metadata to support the provenance such as the date or time, the type of clinical contact, or who assessed the patient etc. We know what was ordered and when, and now we can discern some of the ‘why’ with the benefit of the additional clinical data that is also contained within the report or clinic letter.
The semantic awareness is due to the ability to add additional information to the core message. How useful that will be depends on whether this information is standardised or coded. For example, a CDA document has a specific structure and so a receiving computer should be able to process this. Additionally, a basic HL7 FHIR message also contains semantic data in the way that it requires patient attributes and user viewable HTML section, although these are primarily human readable semantics.
Finally, to make the document useful, we should again make use of a terminology system to provide coded, structured content to enable reuse of the information. The same caveat applies from Use Case 1 in terms of a lack of standardisation, and therefore technical lock-in is still a risk. In some ways this is semantic-lite depending on how well structured the document is.
Use Case 3: Support Clinical Workflow
We are now increasing the reliance on structured data to provide additional cohesion to acquire semantic interoperability. For example, when a discharge advice letter is sent from an acute hospital to the GP, it can be integrated into the software workflows of the recipient system. They then gain semantic understanding of the clinical interactions leading up to the event and are able to use this information directly to support onward care. The GP system knows that there has been a discharge summary, or a diagnostic screening result. Better still, the data conforms to an agreed, standardised information model such as an openEHR archetype.
The recipient system can then automate certain tasks (even if that just means respond to the request to "do something with this"). But this is where the clinical interoperability begins to take hold. We are giving clinical staff the ability to intelligently use data, and for it to inform their decision-making processes, even at a basic level. For this we may need both structured data (openEHR), contained within a standardised message (FHIR) and coded with an effective terminology (Snomed CT).
Use Case 4: Clinical Decision Support
Now clinical/process interoperability gets into the nitty gritty. For example, if a system can discern that an allergic reaction to Substance A should flag a warning if a similar Substance B is prescribed then this may have a direct implication on clinical care. This is where the likes of Snomed CT can really help due to the hierarchal linkages between clinical terms.
Embedding a NICE guideline, for example, now becomes a possibility. But this is getting into very technically difficult terrain as we no longer need to structure the data, but the myriad options that may affect a patient pathway. Thomas Beale put the challenge front and centre in his talk in November 2017 by explaining healthcare is far removed from Amazon sending a parcel to you. At any point a clinical pathway could change and this level of interoperability requires new language and technology to cope with it. So while this is still firmly clinical interoperability, it is more akin to the upper levels of the LCIM.
Why This Matters
Last week saw the HIMSS conference in Las Vegas and interoperability was a hot topic, as evidenced by numerous articles and tweets...
Statutory definition of interoperability in the US #HIMSS18 @INTEROPenAPI @DHCCIO @NHSDigital @AppertaUK @Andy_Kinnear @HealthCION @AdeByrne @SarahFWilkinson @NHSCCIO "interoperability requires legislation because the market failed" ouch! pic.twitter.com/wC5Srdzjq7
— Prof. Joe McDonald (@CompareSoftware) March 7, 2018
Professor McDonald's comment is interesting, although the way that slide is constructed seems to conflate two distinct things: interoperability and open platforms. But legislation to enforce either seems significant.
The extent of the descriptions above are part of the reason why it is hard to describe interoperability clearly. So while I have issues with the wording used by Meaker et al., I can see why the mistake was made. However, I firmly believe that broadening the definition to highlight technical, semantic and clinical facets is vital to avoid misconstruing the scale and difficulty in achieving healthcare interoperability.
This is vitally important when you hear of solutions to interoperability being touted. It would be easy for the Average Joe to think that jumping on board any one of the standards that exist right now would solve all of their informatics woes. If I have a penny for every time I heard "Snomed CT is going to solve our structured data problems" or "FHIR is the answer to interoperability" I could now retire. It is not that simple, but there is a way forward by using all of these technologies to their strengths.
References
- Meaker, R., Bhandal, S., & Roberts, C. M. (2018). Information flow to enable integrated health care: integration or interoperability. Br J Gen Pract, 68(668), 110–111.
- Benson, T., & Grieve, G. (2016). Principles of health interoperability: Snomed CT, HL7 and FHIR. Health Information Technology Standards.
- IEEE. IEEE standard computer dictionary: a compilation of IEEE standard computer glossaries. New York: Institute of Electrical and Electronics Engineers; 1990.
- Tolk, A., Diallo, S. Y., & Turnitsa, C. D. (2007). Applying the Levels of Conceptual Interoperability Model in Support of Integratability, Interoperability, and Composability for System-of-Systems Engineering. Journal of Systemics, Cybernetics and Informatics, 5(5), 65–74. Retrieved from http://www.iiisci.org/journal/cv$/sci/pdfs/p468106.pdf
-
No. I didn’t know what this was. I had to look it up...
↩